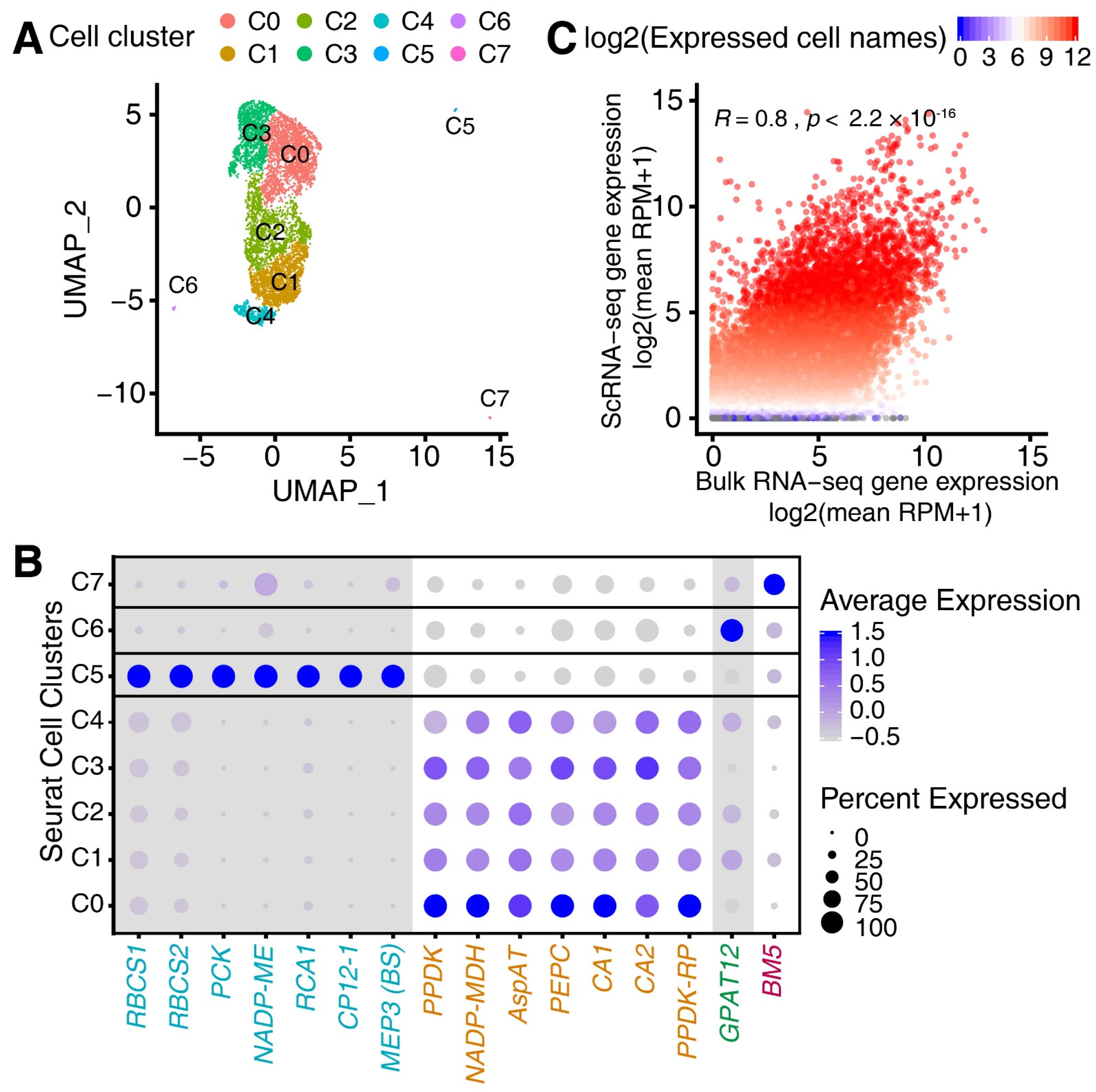
Genes | Free Full-Text | Single-Cell Transcriptome and Network Analyses Unveil Key Transcription Factors Regulating Mesophyll Cell Development in Maize

CReVIS-Seq: A highly accurate and multiplexable method for genome-wide mapping of lentiviral integration sites: Molecular Therapy - Methods & Clinical Development

Dispen-Seq: a single-microparticle dispenser based strategy towards flexible cell barcoding for single-cell RNA sequencing | SpringerLink
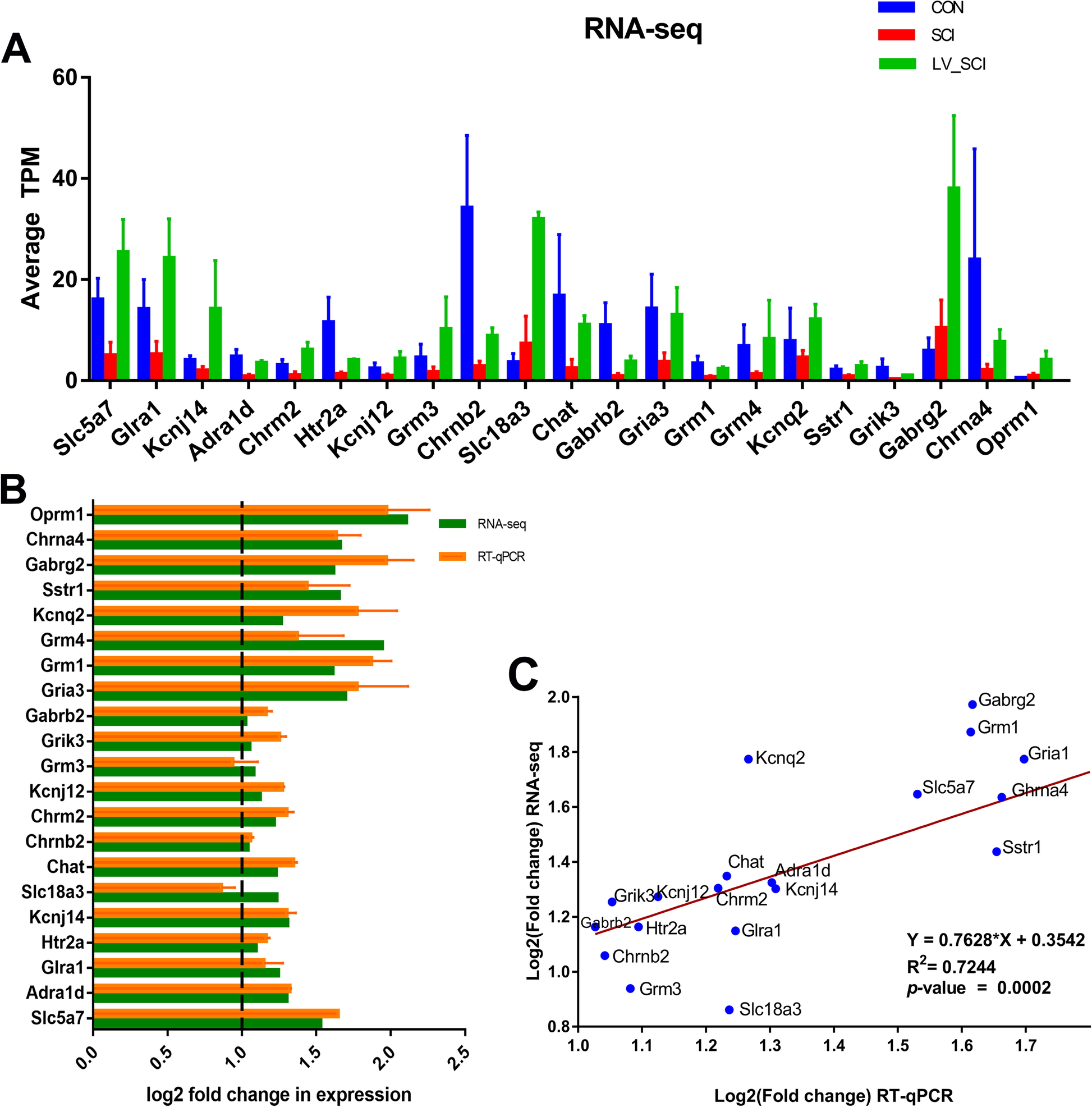
Transcriptomic analysis of α-synuclein knockdown after T3 spinal cord injury in rats | BMC Genomics | Full Text

Slide-seq: A scalable technology for measuring genome-wide expression at high spatial resolution | Science

DNase-seq: A High-Resolution Technique for Mapping Active Gene Regulatory Elements across the Genome from Mammalian Cells
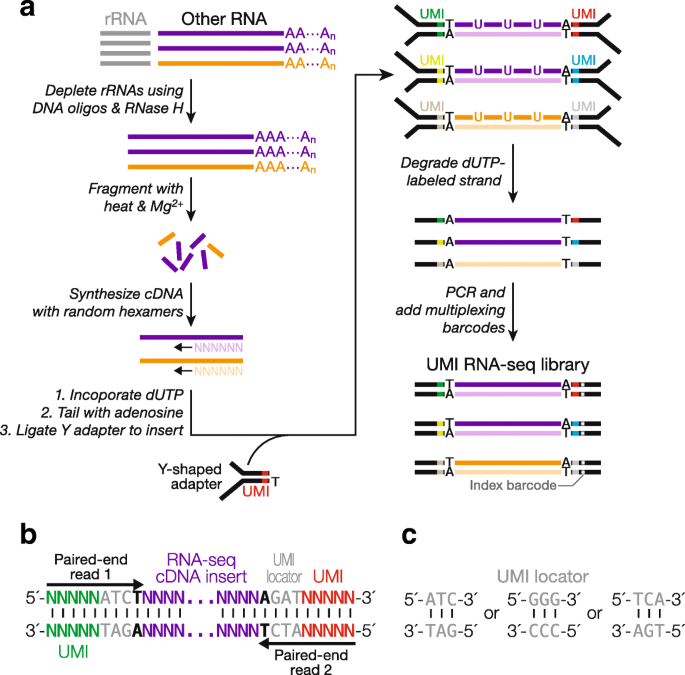
Elimination of PCR duplicates in RNA-seq and small RNA-seq using unique molecular identifiers | BMC Genomics | Full Text

Introducing MAS-ISO-seq: A new long-read sequencing protocol for dramatically increasing RNA transcript isoform yield - Terra.Bio
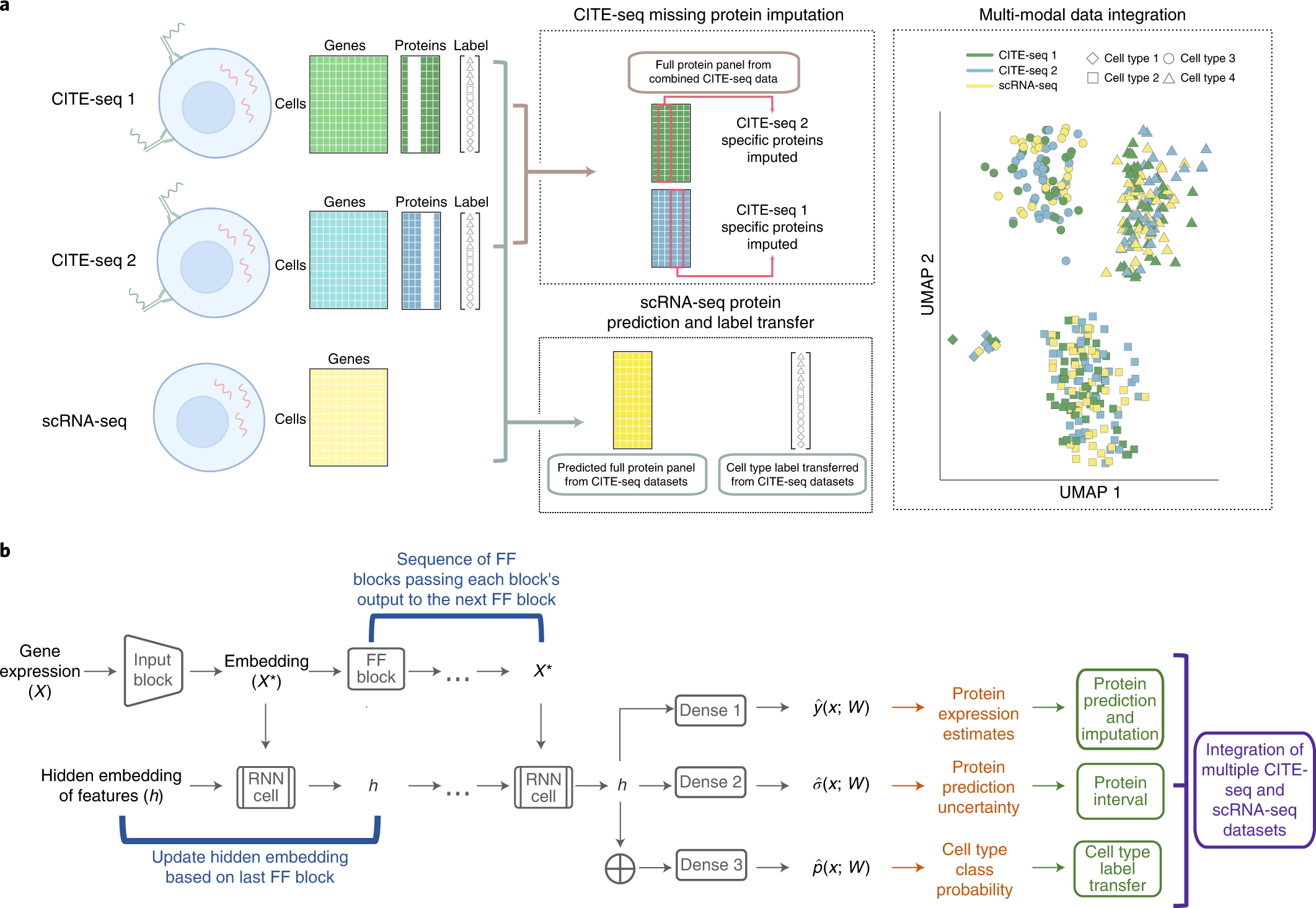
A multi-use deep learning method for CITE-seq and single-cell RNA-seq data integration with cell surface protein prediction and imputation | Nature Machine Intelligence

DiMeLo-seq: a long-read, single-molecule method for mapping protein–DNA interactions genome wide | Nature Methods
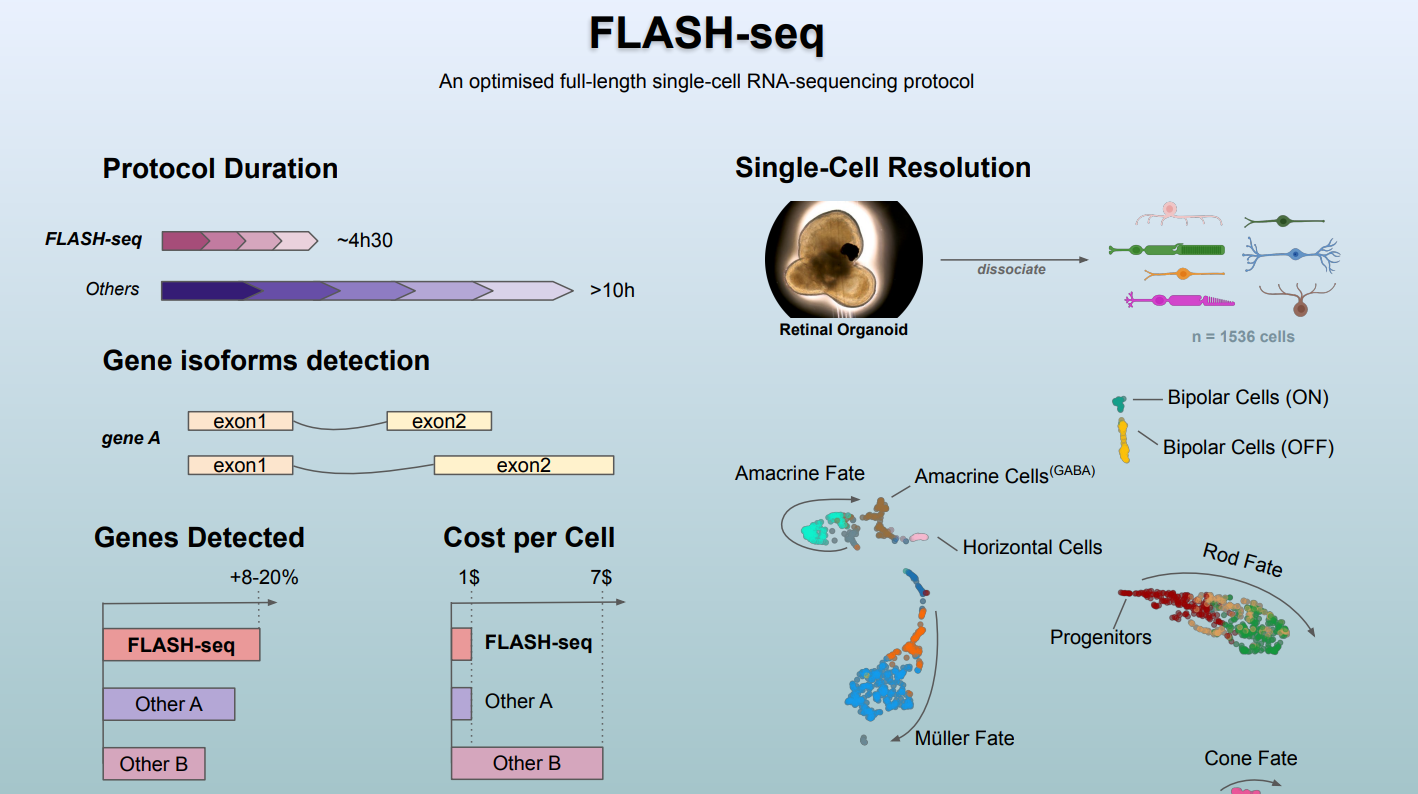
IOB's new scRNA-seq protocol provides a snapshot of the cell transcriptome at unprecedented resolution

3D RNA-seq - a powerful and flexible tool for rapid and accurate differential expression and alternative splicing analysis of RNA-seq data for biologists | bioRxiv

Schematic diagram representing EXPRSS Tag-seq. (A) Library preparation... | Download Scientific Diagram

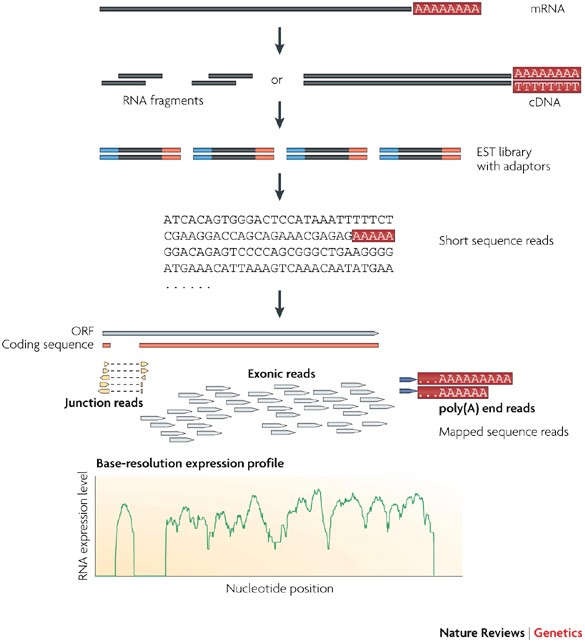
Overview of RNA-seq.A RNA fraction of interest is selected, fragmented... | Download Scientific Diagram